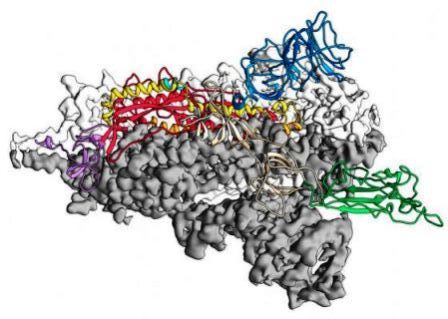
Review on structural mechanism and mode of action of corona virus
Abstract
Human coronaviruses, first reported in the 1960s, were the reason for a large number of children infected with the upper respiratory tract. SARS and four other coronaviruses have been reported since 2003, causing substantial morbidity and mortality. A new coronavirus was firstly reported from Hubei Province, Wuhan City in China, in December 2019 and named as novel Corona Virus. Later on, this virus is referred to as COVID-19, which can cause diseases similar to SARS and named for the spikes that protrude from their membranes, like the sun’s corona. The virus is believed to have been spread by animals sold in Wuhan from the seafood market and Health authorities in China reported fever, cough, breathing problems, and pneumonia in patients. The virus has a variety of physical and pathogenic features similar to the SARS-CoV-2 virus. Closer contact may cause the virus to be transmitted Initial flu-like symptoms, such as fever, cough, myalgia, and dyspnea in the 2-14 days of viral contact. People who are chronically ill are usually older or have medical co-morbidity. No treatments or virus vaccines are currently available. Seasonal Influenza has less mortality rate and less infectious than SARS-CoV-2. Many of the viruses remain to be identified as they will spread. Future directions for SARS-CoV work include greater knowledge of the replication process, tropism, and immune-response mechanisms, taking account of the possible functions of groups-specific proteins; development of animal and human virus vaccine strategies, and anti-viral therapies and very likely isolation and characterization of new human pathogenic coronaviruses. Emergencies for identifying and quarantining contaminated individuals early in the process of preventing their further spread are being made through global efforts
Keywords
Full Text:
PDFReferences
Adhikari, S. P., S. Meng, Y.-J. Wu, Y. P. Mao, R. X. Ye, Q. Z. Wang, C. Sun, S. Sylvia, S. Rozelle, and H. Raat, 2020. Epidemiology, causes, clinical manifestation and diagnosis, prevention, and control of coronavirus disease (covid-19) during the early outbreak period: A scoping review. Infectious diseases of poverty, 9(1): 1-12.
Baseer, M.-A., S.-H. Ansari, S.-S. AlShamrani, A.-R. Alaska's, R. Mahrous, and A.-M. Alenazi, 2016. Awareness of droplet and airborne isolation precautions among dental health professionals during the outbreak of coronavirus infection in Riyadh city, Saudi Arabia. Journal of clinical and experimental dentistry, 8(4): e379.
Corman, V. M., O. Landt, M. Kaiser, R. Molenkamp, A. Meijer, D. K. Chu, T. Bleicker, S. Brünink, J. Schneider and M. L. Schmidt, 2020. Detection of 2019 novel coronavirus (2019-ncov) by real-time RT-PCR. Eurosurveillance, 25(3).1-15
Cui, J., F. Li, and Z.-L. Shi, 2019. Origin and evolution of pathogenic coronaviruses. Nature reviews microbiology, 17(3): 181-192.
Ghinai, I., T. D. McPherson, J. C. Hunter, H. L. Kirking, D. Christiansen, K. Joshi, R. Rubin, S. Morales-Estrada, S. R. Black, and M. Pacelli, 2020. First known person-to-person transmission of severe acute respiratory syndrome coronavirus 2 (sars-cov-2) in the USA. The Lancet.
Hamre, D., and J. J. Procknow, 1966. A new virus is isolated from the human respiratory tract. Proceedings of the society for experimental biology and medicine, 121(1): 190-193.
Hoffmann, M., H. Kleine-Weber, N. Krüger, M. A. Mueller, C. Drosten and S. Pöhlmann, 2020. The novel coronavirus 2019 (2019-ncov) uses the sars-coronavirus receptor ace2 and the cellular protease tmprss2 for entry into target cells. BioRxiv.
Imai, N., I. Dorigatti, A. Cori, C. Donnelly, S. Riley, and N. M. Ferguson, 2020. Report 2: Estimating the potential total number of novel coronavirus cases in Wuhan City, China. Imperial College London.
Jin, Y.-H., L. Cai, Z.-S. Cheng, H. Cheng, T. Deng, Y.-P. Fan, C. Fang, D. Huang, L.-Q. Huang and Q. Huang, 2020. A rapid advice guideline for the diagnosis and treatment of 2019 novel coronavirus (2019-ncov) infected pneumonia (standard version). Military medical research, 7(1): 4.
Lai, C. C., T.-P. Shih, W.-C. Ko, H.J. Tang, and P.R. Hsueh, 2020. Severe acute respiratory syndrome coronavirus 2 (sars-cov-2) and coronavirus disease-2019 (covid-19): The epidemic and the challenges. International journal of antimicrobial agents: 105924.
Lauer, S. A., K. H. Grantz, Q. Bi, F. K. Jones, Q. Zheng, H. R. Meredith, A. S. Azman, N. G. Reich, and J. Lessler, 2020. The incubation period of coronavirus disease 2019 (covid-19) from publicly reported confirmed cases: Estimation and application. Annals of internal medicine. 7(2):11-14.
Li, W., C. Zhang, J. Sui, J. H. Kuhn, M. J. Moore, S. Luo, S. K. Wong, I. C. Huang, K. Xu, and N. Vasilieva, 2005. Receptor and viral determinants of sars‐coronavirus adaptation to human ACE2. The EMBO journal, 24(8): 1634-1643.
Lu, H., 2020. Drug treatment options for the 2019-new coronavirus (2019-ncov). Bioscience trends, 14(1): 69-71.
McIntosh, K., M. S. Hirsch, and A. Bloom, 2020. Coronavirus disease 2019 (covid-19). UpToDate. Hirsch MS, Bloom A (Eds.). Accessed Mar, 5.
Memish, Z. A., A. Zumla, R. F. Alhakeem, A. Assiri, A. Turkestani, K. D. Al Harby, M. Alyemni, K. Dhafar, P. Gautret, and M. Barbeschi, 2014. Hajj: Infectious disease surveillance and control. The Lancet, 383(9934): 2073-2082.
Nishiura, H., 2020. Back calculating the incidence of infection with covid-19 on the diamond princess. Multidisciplinary digital publishing institute.
Nishiura, H., T. Kobayashi, Y. Yang, K. Hayashi, T. Miyama, R. Kinoshita, N. M. Linton, S.-m. Jung, B. Yuan, and A. Suzuki, 2020. The rate of under ascertainment of novel coronavirus (2019-ncov) infection: Estimation using Japanese passenger data on evacuation flights. Multidisciplinary digital publishing institute.
Ontiveros, E., T. S. Kim, T. M. Gallagher, and S. Perlman, 2003. Enhanced virulence mediated by the murine coronavirus, mouse hepatitis virus strain jhm, is associated with glycine at residue 310 of the spike glycoprotein. Journal of virology,
(19): 10260-10269.
Otter, J., C. Donskey, S. Yezli, S. Douthwaite, S. Goldenberg, and D. Weber, 2016. Transmission of sars and mers coronaviruses and influenza virus in healthcare settings: The possible role of dry surface contamination. Journal of hospital infection, 92(3): 235-250.
Paraskevis, D., E. G. Kostaki, G. Magiorkinis, G. Panayiotakopoulos, G. Sourvinos, and S. Tsiodras, 2020. Full-genome evolutionary analysis of the novel coronavirus (2019-ncov) rejects the hypothesis of emergence as a result of a recent recombination event. Infection, genetics, and evolution, 79: 104212.
Park, S. H., Y. S. Kim, Y. Jung, N. H. Cho, H. W. Jeong, J. Y. Heo, J. H. Yoon, J. Lee, S. Cheon, and K. M. Sohn, 2016. Outbreaks of middle east respiratory syndrome in two hospitals initiated by a single patient in Daejeon, South Korea. Infection & chemotherapy, 48(2): 99-107.
Rothe, C., M. Schunk, P. Sothmann, G. Bretzel, G. Froeschl, C. Wallrauch, T. Zimmer, V. Thiel, C. Janke, and W. Guggemos, 2020. Transmission of 2019-ncov infection from an asymptomatic contact in Germany. New England Journal of Medicine. 7(3):11-16.
Sahin, A. R., A. Erdogan, P. M. Agaoglu, Y. Dineri, A. Y. Cakirci, M. E. Senel, R. A. Okyay, and A. M. Tasdogan, 2020. 2019 novel coronavirus (covid-19) outbreak: A review of the current literature. The EJMO, 4(1): 1-7.
Song, Z., Y. Xu, L. Bao, L. Zhang, P. Yu, Y. Qu, H. Zhu, W. Zhao, Y. Han, and C. Qin, 2019. From sars to mers, thrusting coronaviruses into the spotlight. Viruses, 11(1): 59.
Syed, A., 2020. Coronavirus: A mini-review. International journal of current research and medicine sciences, 6(1): 8-10.
Tyrrell, D., J. Almeida, C. Cunningham, W. Dowdle, M. Hofstad, K. McIntosh, M. Tajima, L. Y. Zakstelskaya, B. Easterday, and A. Kapikian, 1975. Coronaviridae. Intervirology, 5(1-2): 76-82.
Ullah, E., 2020. The novel coronavirus outbreak: A challenge beyond borders. Pakistan journal of sugar medicane, 1(1): 8-9.
Wang, W., X.-D. Lin, W. P. Guo, R. H. Zhou, M.-R. Wang, C. Q. Wang, S. Ge, S. H. Mei, M. H. Li, and M. Shi, 2015. Discovery, diversity, and evolution of novel coronaviruses sampled from rodents in China. Virology, 474: 19-27.
Woo, P. C., Y. Huang, S. K. Lau, and K.-Y. Yuen, 2010. Coronavirus genomics and bioinformatics analysis. Viruses, 2(8): 1804-1820.
Wrapp, D., N. Wang, K. S. Corbett, J. A. Goldsmith, C.L. Hsieh, O. Abiona, B. S. Graham, and J. S. McLellan, 2020. Cryo-em structure of the 2019-ncov spike in the prefusion conformation. Science, 367(6483): 1260-1263.
Wu, F., S. Zhao, B. Yu, Y.-M. Chen, W. Wang, Z.-G. Song, Y. Hu, Z.-W. Tao, J. H. Tian, and Y. Y. Pei, 2020. A new coronavirus is associated with human respiratory disease in China. Nature, 579(7798): 265-269.
Xu, X., P. Chen, J. Wang, J. Feng, H. Zhou, X. Li, W. Zhong, and P. Hao, 2020. Evolution of the novel coronavirus from the ongoing Wuhan outbreak and modeling of its spike protein for risk of human transmission. Science China life sciences, 63(3): 457-460.
Yin, Y., and R. G. Wunderink, 2018. Mers, sars, and other coronaviruses as causes of pneumonia. Respirology, 23(2): 130-137.
Zhang, M.-J., D.-J. Liu, X.-L. Liu, X.-Y. Ge, A. Jongkaewwattana, Q. G. He, and R. Luo, 2019. Genomic characterization and pathogenicity of porcine delta coronavirus strain chn-hg-2017 from China. Archives of virology, 164(2): 413-425.
Zhu, N., D. Zhang, W. Wang, X. Li, B. Yang, J. Song, X. Zhao, B. Huang, W. Shi, and R. Lu, 2020. A novel coronavirus from patients with pneumonia in china, 2019. New England journal of medicine. 382(8): 727-733
_______________________________________________________
Abstract Views or DF Downloads
DOI: https://doi.org/10.33865/wjb.005.03.0377
Refbacks
- There are currently no refbacks.
Copyright (c) 2020 Muhammad Talha, Hafiz Ghulam Muhu-Din Ahmed, Aziz Ullah, Muhammad Ali
License URL: https://creativecommons.org/licenses/by/4.0/
Print ISSN: 2522-6746 : Online ISSN: 2522-6754
The journal World Journal of Biology and Biotechnology has migrated to a new website after Volume 10 (10 years), and all new submissions and publications are handled through the updated official website.”
https://journals.sciplatform.com/index.php/wjb